A new platform for brain-wide imaging and reconstruction of neurons
The fine, branching tendrils that extend from a neuron often traverse great distances to form connections with their target cells. Tracing those connections is a major goal for scientists who want to understand exactly how the brain processes information, but technical obstacles have hindered the effort. Now, scientists at the Howard Hughes Medical Institute's Janelia Research Campus have traced the complete paths of several neurons through the entire brain of a mouse, using technology they say can be scaled up to enable a larger mapping effort.
A multidisciplinary project team of Janelia scientists reports the development of a new imaging platform that combines a high-speed, high-resolution light microscope with new sample preparation methods and powerful image processing software to efficiently map neuronal projections cell by cell. They published their research online on January 20, 2016 in the journal eLife. "Being able to identify the targets of single neurons has been very difficult to do in the past," says Michael Economo, a research scientist at Janelia who is the first author of the publication. "This microscope should really make that process more efficient."
The publication includes reconstructions of five neurons that spread across the mouse brain in complex and diverse patterns, representing a milestone for Janelia's MouseLight Project team, which aims to generate maps of neurons' projections throughout the mouse brain.
"The goal of the project is to see where individual neurons in the brain send their messages, and to do this at scale," says project leader Jayaram Chandrashekar. Ultimately, he says, the MouseLight team aims to trace the projections of thousands of neurons, a small percentage of the 70 million neurons in the mouse brain, but a huge increase over the handful of projections that have been reconstructed to date. "To do this," he says, "we need to be able to follow individual neurons right from their cell body all the way to their termini, so we need very good resolution, and we need speed. The current technology lets us do both of these things."
Although scientists are beginning to map out how different areas of the mouse brain connect to one another, they have little understanding of the cell-to-cell variation that exists within these broader connections. "There's a major mystery about how many projection types there are, and this problem has been addressed in a very piecemeal manner," says Karel Svoboda, a Janelia group leader who helped guide the development of the MouseLight project.
By tracing the projections of a significant percentage of the cells in the brain, the team expects to learn a lot about how the nervous system processes information. "A neuron's projection field is complex but highly specific, and it essentially defines where that neuron sends information," Svoboda says. "So knowing a neuron's shape in great detail tells us about how signals are routed in the brain."
The neuron-mapping effort began in the lab of former Janelia group leader Gene Myers, a computer scientist who is now at the Max Planck Institute of Molecular Cell Biology and Genetics. In 2010, Nathan Clack, a scientist in Myers's lab (now at Vidrio Technologies), set out to apply imaging analysis methods being developed in Myers's lab to the problem of tracing neurons in the mouse brain.
"It's very difficult to take a picture of an entire neuron," Clack says. "The axon—the business end of the neuron—is like a thin wire, as little as 80 nanometers in diameter, that travels for centimeters." To capture a neuron's slender extensions from beginning to end, he says "you really have to think about imaging the entire brain."
Clack's first priority, then, was to optimize a high-resolution microscope for the task. Working with Janelia's Instrument Design & Fabrication team, he adapted a two-photon microscope—a technology that lets scientists image clearly beneath the surface of a tissue—to speed up its imaging rate dramatically. Even with the improved speed, the microscope would still take more than a week to image a complete mouse brain, so Clack developed control software to fully automate the process.
Clack then teamed with Chandrashekar and Economo to work out a method of readying a mouse brain for imaging. Brain tissue is clouded with fats that obscure the view beneath the surface, so unless these fats are removed, the tissue must be sliced into ultrathin sections to obtain high-resolution images. That process that can distort tissue, making it difficult to reconstruct images of structures that pass through many sections. Existing methods of removing fats were not compatible with a prolonged imaging period, so Janelia group leader Luke Lavis, a chemist, applied his expertise to the problem. Ultimately, the team developed a mild clearing strategy that allows the microscope to image clearly to about 250 microns beneath the surface of a sample. "This approach preserves the fluorescence [in labeled neurons] and gives us really high quality data," Clack says.
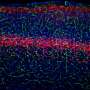
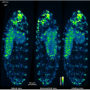
To generate the images they would use to trace neuronal projections, Economo and his colleagues applied their clearing protocol to a mouse brain in which a small number of neurons had been brightly labeled with a fluorescent dye. They inserted this brain into the microscope and, after some initial set-up, automated imaging began.
To image a complete brain, the microscope collects about 20,000 separate but overlapping blocks of images, imaging as deep as it can, then slicing away a layer of tissue and repeating the process. The entire brain is imaged in about 100 slices, a process that takes about 7-10 days and generates tens of terabytes of data. Taking advantage of methods developed by Janelia's scientific computing group to manage the large volume of data, the 20,000 image blocks are ultimately computationally stitched together to form a complete three-dimensional representation of the brain.
The last step, tracing the paths of the fluorescently labeled neurons to generate final reconstructions of each cell's projections, was done manually, demanding 40 to 60 hours of work per cell. The team is now exploring ways to dramatically reduce the time that must be devoted to this task, working to improve cell labeling and computational analysis so that tracing can be done mostly automatically. "This step will never be 100 percent automated, but we believe it can be brought down to a few hours," Chandrashekar says. That will be essential as the team works toward using their platform to complete about one neuronal reconstruction per day.