New technique helps researchers determine developmental origins of cells
Howard Hughes Medical Institute (HHMI) scientists have pioneered the use of genome editing to trace lineage in living systems. The researchers introduced unique patterns of mutations into the cells of a developing zebrafish embryo using the CRISPR/Cas9 system. These mutations persist into adulthood and act as barcodes that scientists can read to reveal the relationships between individual cells during development. The method will help researchers map the complex series of cell divisions that transforms a single cell into a complete organism.
"You could call this a shotgun approach to developmental biology," says Jay Shendure, an HHMI investigator at the University of Washington, who co-led the development of the new technique with collaborator Alexander Schier at Harvard University. "We're packing a lot of information about lineage into a very short region [of DNA] that we can then sequence very efficiently." That efficiency means researchers can use the barcodes to determine the lineages of large numbers of cells without perturbing their normal developmental path.
In a paper published May 27, 2016, in the journal Science, Shendure and colleagues report that they have used the method, which they call GESTALT (Genome Editing of Synthetic Target Arrays for Lineage Tracing), to determine the developmental origins of hundreds of thousands of cells in the bodies of adult zebrafish. The three co-first authors of the work are Aaron McKenna and Greg Findlay, graduate students at the University of Washington, and Jamie Gagnon, a postdoctoral fellow at Harvard University.
Every multicellular organism develops from a single fertilized egg. As cells in the embryo divide continuously, information encoded in the genome of those cells directs their specialization into various types of tissues and organs. Mapping out these cell divisions and the fates of the cells they produce has long been a goal of developmental biologists, but so far this has only been achieved on a cell-by-cell basis for the roundworm Caenorhabditis elegans. C. elegans has only about a thousand cells when fully developed. The task is far more challenging in larger, more complex organisms.
One strategy for tracing cell lineages in more complex organism—where cell divisions cannot be followed visually—is to identify and track in adult cells the mutations that arose early in development. Those mutations are passed along to new cells with each cell division, so cells with shared ancestry will end up with some of the same mutations. But these naturally occurring mutations can arise anywhere in a cell's DNA, and researchers must completely sequence the genomes of individual cells to find them. This approach is currently too costly to use for all the cells of an adult organism.
Shendure and his colleagues thought they could solve this problem by creating a compact segment of DNA where mutations could accumulate during development. Later, they could sequence just that bit of DNA, and each cell's mutations would serve as a record of its developmental path.
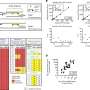
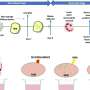
To ensure that this piece of DNA would accumulate enough mutations to identify each cell's lineage, the researchers designed it to be targeted by a genome editing system, CRISPR/Cas9. Researchers use CRISPR/Cas9 to make changes at specific locations in a DNA molecule. Shendure and his colleagues designed their short stretch of DNA to be packed with up to 12 different CRISPR/Cas9 target sites.
They introduced these genetic barcodes into cells, and then used the CRISPR/Cas9 system to create mutations at the target sites. They tested the approach first in lab-grown cells, and then in a zebrafish embryo. Barcode editing continued for a few hours after the CRISPR/Cas9 tools were introduced, so that dividing cells acquired new mutations as time went by.
Shendure and his colleagues could then examine the barcodes inside individual cells and look for shared mutations. They knew that the more mutations two cells shared, the more recently they had diverged from the same progenitor cell. They used that information to reconstruct cell lineages.
Some mutations, or edits, were present in half of the cells that the researchers analyzed—suggesting that the CRISPR system had made those changes when the embryo consisted of just two cells. "As more and more cell divisions occur, you get the accumulation of more and more edits that are unique to a sub-lineage, or a sub-sub-lineage, or a sub-sub-sub-lineage," Shendure explains.
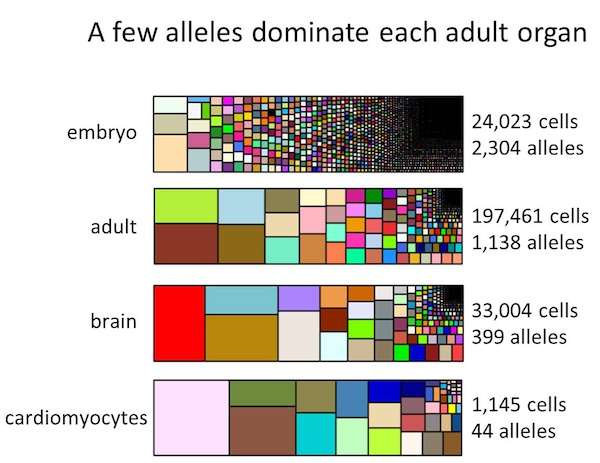
The team analyzed about 200,000 cells from each of two adult zebrafish, taken from many different tissues, including the heart, eyes, blood, and brain. Using the barcode information, they mapped out how different cells in the embryo contribute to different adult organs. In doing so, they found that the majority of cells sampled from specific organs descended from just a few embryonic progenitors.
An important next step, Shendure says, will be developing a way to link information about each cell's lineage to additional information about the same cell, such as which genes it has turned on or its exact position within a tissue. "Then we can get much more fine-grained about what we can say about developmental biology in terms where specific cell types come from and how they're related to each other," he says.
More information: A. McKenna et al. Whole organism lineage tracing by combinatorial and cumulative genome editing, Science (2016). DOI: 10.1126/science.aaf7907