Scientists find gene linked to heightened mucus levels in lung disease
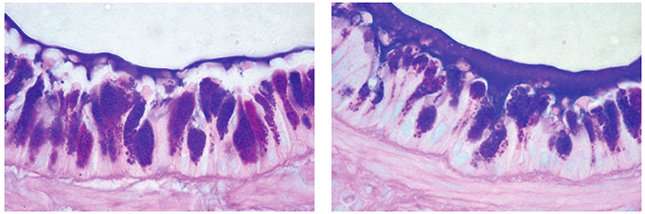
What if researchers could make breathing easier by changing how much mucus is in your lungs? Although healthy individuals have mucus in their lungs, mucus can be a major problem for people with chronic airway conditions, such as asthma, cystic fibrosis (CF) and chronic obstructive pulmonary disease (COPD). In a paper published in Genetics, UNC School of Medicine researchers led by Samir Kelada, PhD, MPH, assistant professor of genetics, showed that the gene Bpifb1 is strongly associated with one of the major mucin proteins (MUC5B) that controls the physical nature of mucus in the lungs.
Lung mucus has important roles, including trapping infections. But it's important to not have too much mucus. If there's too much mucus or it's too sticky, then it can build up in the lung, blocking the space for the air we breathe. This can lead to difficulty breathing or increased risk of infections, making diseases or conditions like asthma worse. Tweaking the level of mucin proteins such as MUC5B could be a key part of therapies.
Using the Collaborative Cross – populations of laboratory mice derived from mixing eight genetically diverse inbred lines – Kelada and colleagues were able to map a genomic location that controls mucin levels, and this region contains the gene Bpifb1. Once they accomplished that, they looked at what happened when the gene was deleted and found a three-fold increase of MUC5B levels in the airways of mice.
"Other researchers at the UNC Marsico Lung Institute previously found that even a seemingly slight increase of mucins can lead to severe changes to mucus in human airways," said Kelada, a Marsico member. "Our finding shows that Bpifb1 plays a causative role in MUC5B levels in the airways. How exactly it does this is still not known, but our work suggests the gene is involved in regulating biophysical aspects of mucus, and this regulation is crucial to how well a person clears mucus."
Although this work was done in mice, humans have the Bpifb1 gene. Thus, the UNC researchers will now explore whether genetic variation in Bpifb1 can explain differences in mucus accumulation in humans with asthma, CF, or COPD.
"Our paper also highlights the great utility of the Collaborative Cross in mapping genes, like Bpifb1, for biomedically relevant traits," Kelada said. "In collaboration with investigators at the Marsico Lung Institute, we will be moving beyond gene discovery to using the Bpifb1 knockout mouse to explore fundamental aspects of mucus structure and function. Now the goal is to find ways to alter mucus so it becomes much less of a problem for people with these conditions."
More information: Lauren J. Donoghue et al. Identification of trans Protein QTL for Secreted Airway Mucins in Mice and a Causal Role for Bpifb1, Genetics (2017). DOI: 10.1534/genetics.117.300211