Newly identified subset of immune cells may be key to fighting chronic inflammation

St. Jude Children's Research Hospital scientists have discovered a subset of helper T cells that may help to redefine understanding and treatment of chronic, debilitating inflammatory disorders. The research appears today as an advance online publication in the journal Nature.
The study focused on a family of helper T cells called Th17 cells. Th17 cells help to launch the immune response against fungal infection and other threats. These cells can also fuel the destructive inflammation in autoimmune disorders like multiple sclerosis and rheumatoid arthritis.
Th17 cells have been known for their functional adaptability (plasticity), but the mechanism that helped to regulate plasticity was unclear.
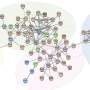
Now investigators have identified a new subset of Th17 cells that is functionally and metabolically distinct from Th17 cells associated with chronic inflammation. Researchers reported that the two Th17 cell subsets expressed unique transcription factors that regulate activity of different genes. The scientists showed that gene expression, and thus cell function, was regulated in part by metabolism.
"These results provide a missing link that helps explain Th17 plasticity, including the cellular process that explains how cell function changes and the role metabolism plays in regulating the process," said corresponding author Hongbo Chi, Ph.D., a member of the St. Jude Department of Immunology. "Identification of this new Th17 subset opens new avenues for developing effective treatment of chronic inflammatory conditions."
Metabolic activity regulates gene expression
The research builds on previous findings from Chi and his colleagues. Those studies highlighted the protein complex mTORC1 and metabolic activity as important regulators of activation and function of T cells. mTORC1 helps regulate cell growth and metabolism by coordinating cell metabolism and transcription.
In this study, researchers used a mouse model of human multiple sclerosis and the tools of systems biology, including single cell RNA sequencing and genome-wide chromatin accessibility assays, to show that metabolic activity and mTORC1 play important roles in regulating Th17 function.
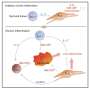
Investigators found that disrupting either mTORC1 function or reducing metabolic activity in Th17 cells shifted gene expression to favor expression of the transcription factor TCF-1. TCF-1 turns on genes associated with blood stem cells. These are "immature" cells with the ability to differentiate into cells with more specific functions. Increased TCF-1 expression led to Th17 cells that were more "stem-like" and quiescent.
In contrast, higher metabolic activity in Th17 cells favored expression of a different transcription factor, T-bet. T-bet is associated with elevated levels of interferon gamma, a molecule that fuels inflammation and expressed by another type of helper T cells, Th1 cells.
Researchers have known that Th17 cells that function like Th1 cells are a hallmark of chronic inflammatory disorders like multiple sclerosis. This study found that disrupting either mTORC1 or metabolic activity blocked Th1-like features in Th17 cells in the laboratory as well as neuroinflammation in mice.
Clinical hopes
Research is underway to determine if the newly identified subset of Th17 cells, those with TCF-1 expression and stem-like features, fuels inflammatory diseases by serving as a reservoir of cells that can be used to resupply inflammation-promoting Th17 cells. "The discovery of a pool of stem-like Th17 cells may open new therapeutic avenues for treatment of chronic inflammatory diseases," Chi said.
More information: Peer W. F. Karmaus et al. Metabolic heterogeneity underlies reciprocal fates of TH17 cell stemness and plasticity, Nature (2018). DOI: 10.1038/s41586-018-0806-7