Researchers speed up detection of blood infection
Researchers from The University of Western Australia have developed a new method of detecting blood infection that they hope will dramatically speed up diagnosis and treatment of severe infection.
The research follows last year's exciting results from a screening test to detect antibiotic resistance and ensure the right antibiotics can be prescribed quicker.
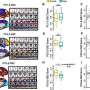
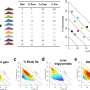
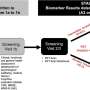
The time-saving solution known as FISH-flow (since it combines Fluorescent In Situ Hybridisation with flow cytometry) accurately detects bacteria in blood hours before current routine methods can.
Flow cytometry is a powerful laboratory tool that uses lasers, digital electronics and graphic imaging technology to analyse the physical and chemical characteristics of single cells in a liquid, such as blood or bone marrow.
Professor Tim Inglis, Head of Pathology and Laboratory Medicine at UWA, said the significance of the new method was that it would complement the ultra-rapid method of antibiotic testing his group is working on, to speed up diagnosis and treatment of serious bloodstream infections.
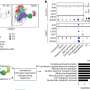
Professor Inglis said the research paper, published in PLOS One, described the application of flow cytometry to early detection of bacteria in blood so that laboratory diagnosis of bloodstream infection could be accelerated.
He said any reduction in the time it took to choose the right antibiotic was likely to save lives.
Bloodstream infections, often known as sepsis, remain a common cause of serious illness and death in developed countries, and are becoming increasingly difficult to treat effectively due to rising levels of antibiotic resistance.
More information: Xiao Xuan Huang et al. Accelerated bacterial detection in blood culture by enhanced acoustic flow cytometry (AFC) following peptide nucleic acid fluorescence in situ hybridization (PNA-FISH), PLOS ONE (2019). DOI: 10.1371/journal.pone.0201332