Researchers discover how three-dimensional organization of the genome regulates cell differentiation
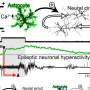
A new study from the University of Minnesota Medical School clarifies how the three-dimensional organization of the genome is regulated at the onset of skeletal muscle formation. Although the DNA sequence of the genome is a linear code, like a long sentence, the actual DNA molecule twists and folds in 3-D space, with some sequences that are distant from each other in sequence being physically close to each other in space. These 3-D interactions are thought to allow proteins that bind DNA to regulate the activity of genes that are distant from where they are binding.
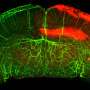
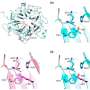
In an article just published in Nature Communications, Alessandro Magli, Ph.D., Assistant Professor, and Rita Perlingeiro, Ph.D., Professor of Medicine and Lillehei Professor in Stem Cell and Regenerative Cardiovascular Medicine, Cardiovascular Division at the University of Minnesota Medical School, and colleagues study the activity of one of these DNA-binding proteins, Pax3, a protein that is essential for the development of skeletal muscle. They show that large loops form between DNA sequences where Pax3 binds, and muscle-specific genes that Pax3 regulates, and that are important for the development of muscle.
The same research group had previously mapped all of the sites to which Pax3 binds in the genome of mesoderm cells, a type of embryonic tissue with the capacity to form muscle. They found that when Pax3 binds to DNA, it opens up and loosens the packing of the DNA molecule in the local region. In the current study, they discovered that after this happens, the Pax3 protein brings in a bridging molecule, called Ldb1, which allows 3 dimensional loops to form between distant sequences by forming a bridge between the Pax3 protein and the distant gene that it is regulating.
To prove the requirement of Ldb1 for the changes in genome architecture, Magli said "they used a genetic trick," as they forced Ldb1 recruitment where Pax3 typically binds in the genome by creating a Pax3-Ldb1 chimera. They also showed that in the absence of Ldb1, muscle does not form properly in mouse embryos.
"Altogether, we demonstrate that Pax3, besides regulating the chromatin architecture, is also manipulating the activation of myogenic enhancers, genomic DNA fragments with important regulatory functions for gene expression," said Magli.
There is evidence that changes in DNA molecular looping, also called chromatin architecture, can lead to genetic disorders and cancer. "By studying how chromatin architecture is regulated within the nucleus, we will better understand both normal and pathological processes and potentially identify new targets for the treatment of diseases such as cancer," said Magli.
"Both publications are great examples of how pluripotent stem cells represent a valuable tool to study cell fate specification and dissect transcriptional regulation," says Perlingeiro.
There is more work ahead, and more questions to be answered. One of Magli's goals is to continue this line of research by determining the signals that induces changes in chromatin architecture and use this knowledge to efficiently generate cells for skeletal muscle regeneration in diseases like muscular dystrophy.
More information: Alessandro Magli et al. Pax3 cooperates with Ldb1 to direct local chromosome architecture during myogenic lineage specification, Nature Communications (2019). DOI: 10.1038/s41467-019-10318-6