A better early blood test for autism: Genetic signatures point to disrupted neuro-immune pathways

(Medical Xpress)—Researchers at Boston Children's Hospital have developed a blood test for autism spectrum disorders (ASDs) that outperforms existing genetic tests, while presenting evidence that abnormal immunologic activity affecting brain development may help explain some of autism's origins. The findings also suggest a new direction for genetic research on autism and the search for treatments.
The blood test, described December 5 in the online open access journal PLOS ONE and based on the largest gene-chip investigation ever done in autism, could enable early diagnosis of autism in about two thirds of patients before clear symptoms start to appear (the average age of diagnosis in the U.S. is 5 years).
Researchers led by Sek Won Kong, MD, of the Boston Children's Hospital Informatics Program (CHIP) analyzed blood samples from 66 male patients with ASDs (from Boston Children's and several other hospitals in collaboration with the Autism Consortium of Boston) and compared them with 33 age-matched boys without ASDs. Using microarrays, they looked for RNA signatures reflecting differences in gene activity, or expression, between the two groups.
"Since brain biopsy isn't a viable option for research, we asked whether blood could serve as a proxy for gene expression in the brain," says Isaac Kohane, MD, PhD, director of CHIP and senior investigator on both studies. "We found that it could, though we and others were initially skeptical."
Analyzing the blood samples, Kong and colleagues flagged 489 genes as having distinct expression patterns in the ASD group, then narrowed this to a group of 55 genes that correctly identified or ruled out autism in 76 percent of samples. They validated their findings in a second group of 104 male and female patients with ASDs and 82 controls, achieving an overall classification accuracy of 68 percent (73 percent for males and 64 percent for females).
The gene signature approach, which Boston Children's Hospital has licensed exclusively to SynapDx (Southborough, Mass.), can potentially diagnose autism far more often than the genetic tests currently available. Those tests look forvariety of autism-related mutations—from small "spelling" changes to lost or extra copies of a gene or genes (known as copy number variants) to wholesale chromosome abnormalities—but together, the known mutations account for fewer than 20 percent of autism cases.

"It's clear that no single mutation or even a single pathway is responsible for all cases," says Kohane. "By looking at this 55-gene signature, which can capture disruptions in multiple pathways at once, we can say with about 70 percent accuracy, 'this child does not have autism,' or 'this child could be at risk,' putting him at the head of the queue for early intervention and evaluation. And we can do it relatively inexpensively and quickly."
Previous gene-expression studies on blood samples have been limited by study size and by the challenge of obtaining well-matched control groups, Kohane notes.
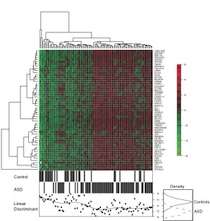
The 55 genes whose expression was altered also suggest more than one path to what we know as autism. Based on their genetic signatures, subjects with ASDs clustered into four subgroups marked by changes in different biological pathways (see image 2):
- Synaptic pathways, specifically long-term potentiation pathways (essential for memory and learning) and neurotrophic pathways (signaling neurons to survive, develop and grow)
- Immune/inflammatory pathways
Adding to the neuro-immune theory of autism
Most current theories of autism focus on disordered synapses (connections between brain cells). But, citingprevious studies in the medical literature, Kohane speculates that brain development in autism may be impaired by abnormal immune responses to infections and other stressors, during infancy or prenatally. For example, chemokines (inflammatory chemical messengers that recruit immune cells to fight infections) have been found to be overexpressed in some neurocognitive disorders, and animal studies have found that prenatal infections are associated with autism-like features in offspring.
"We know that if a mother has rheumatoid arthritis or the father has type 1 diabetes, there is an increased risk of autism in the child," says Kohane. "Studies have also noted an increase in autoimmune disease, just as autism is increasing."
Previous studies of brain tissue from patients with autism have found significant activation of immune cells in the cerebral cortex and cerebellum, and recent research at Boston Children's Hospital has shown that the same molecules used by the immune system also play a role in the development and functioning of the brain.
"One can imagine that an infectious assault during pregnancy could increase chemokines into the brain," Kohane says. "Or there may be something intrinsic to the brain that causes it to generate an inflammatory reaction. Genetic makeup may set up vulnerability in a subgroup of infants, but there may be some environmental factor that triggers the disease in that subgroup."
More information:
Ashwood P; et al. J Leukoc Biol 2006 Jul; 80(1):1-15.
Hsiao EY; et al. Proc Natl Acad Sci U S A 2012 Jul 31; 109(31):12776-81
Atladottir HO; et al. Pediatrics 2009 Aug; 124(2):687-94
Voineagu I; et al. Nature 2011 May 25; 474(7351):380-4
Schafer DP; et al. Neuron 2012 May 24; 74(4):691-705